Highlight
Dynamic Flux Balance Analysis for Mixed-Culture Fermentation
Achievement/Results
For the production of microbial biofuels from lignocellulosic biomass to be economically viable, the fermentable sugars produced from biomass hydrolysis must be utilized efficiently. Most research in this area focuses on creating an omnipotent microorganism that can ferment all sugars to the desired product using genetic engineering techniques. When grown on mixed sugars these microbes exhibit diauxic growth, wherein a more favorable substrate represses the uptake of other carbon sources. Engineering a consortium of specialized microorganisms offers an alternative to the single cell approach that avoids diauxic growth. The objective of this approach is to exploit and enhance the innate capabilities of each cell type in the consortium instead of introducing nonnative functionality.
Timothy Hanly, a Chemical Engineering IGERT trainee in the Henson laboratory at the University of Massachusetts Amherst, is applying metabolic modeling to the study of defined microbial consortia. The metabolic pathways of many microorganisms of interest to research and industry have been mathematically reconstructed. These reconstructions have been used to model steady state cell growth and product formation rates when a cell is subjected to varied culture conditions and genetic engineering. Dynamic flux balance analysis introduces substrate uptake kinetics and extracellular mass balances in order to model transient growth phenomena in batch and fed-batch culture prevalent in industry.
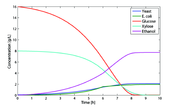
In a study published as the February Spotlight feature in Biotechnology and Bioengineering, Hanly and Henson have combined single cell genome scale flux balance models to simulate a co-culture of Saccharomyces cerevisiae and a mutant strain of Escherichia coli growing on a mixture of glucose and xylose. The microbes in this consortium are specialized and will not compete for the substrates, as S. cerevisiae can only utilize glucose, while the gene deletions in the E. coli mutant ZSC113 have rendered it unable to consume glucose. The dynamic flux balance simulation of the consortium that yielded the highest ethanol productivity at the conditions studied is shown in Figure 1. Substrate uptake parameters for each cell type were found at a common, optimal temperature and pH using pure cultures of each microbe. These parameters were assumed to be constant during mixed culture growth.
A Coulter counter that was previously purchased with IGERT funds and located in the Roberts lab at the University of Massachusetts Amherst was used to distinguish the two microorganisms growing in co-culture. Growth of the mixed culture in a bioreactor was compared to the model predictions. The inhibitory effect of ethanol produced by S. cerevisiae on E. coli growth was identified as the major interaction between the microbes that affected cell metabolism. The model was used to determine the initial amounts of each cell type that would result in simultaneous sugar exhaustion for a variety of glucose and xylose concentrations. Co-cultures of the predicted inocula were implemented in vitro and showed very good agreement with the model predictions.
Address Goals
The complexity of microbial consortia is an obstacle to the application of these systems in industry. The procedure used to construct and validate this model can be applied to other co-cultures of microorganisms for which genome-scale reconstructions are available. A well-defined mixed culture model will be able to predict changes to both intracellular and intercellular dynamics when the co-culture is perturbed. The application of flux balance analysis to mixed culture biotechnology will help expedite research and development of these systems for the production of renewable chemicals. Furthermore, models like the one developed in this work will allow the in silico identification of promising genetic and process engineering strategies to be applied in vitro.